The predictability of evolution has long been a captivating question in evolutionary biology. Because natural selection comprises deterministic components, the course of evolution may exhibit some level of predictability across organismal groups. But just how accurately might we predict evolutionary change?
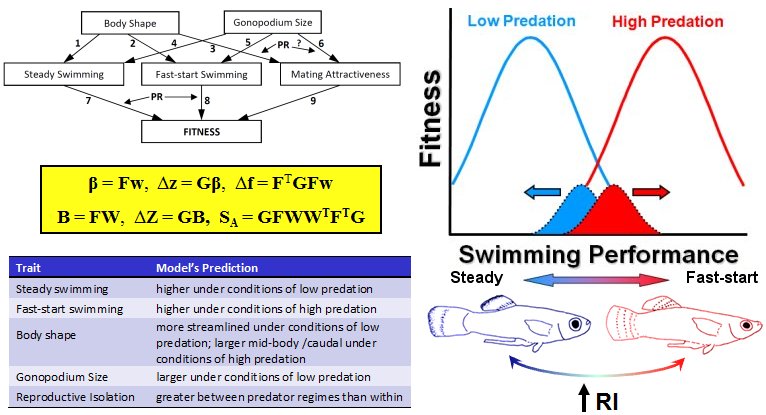
Today, the science of evolutionary prediction is still in its infancy, lacking a cohesive conceptual framework. We have recently proposed, and are continuing to develop, one particular approach to understanding the predictability of evolution: generalized models of divergent selection (GMDS). The GMDS approach is meant to provide a unifying framework for the science of evolutionary prediction, offering a means of better understanding the causes and consequences of phenotypic and genetic evolution. Information to the right illustrates the GMDS approach (see Langerhans in press).
Major Goals Addressed by the GMDS Approach:
1. assess predictability and peculiarity of phenotypic / genetic change
2. identify selective agents responsible for major evolutionary patterns
3. reveal mechanisms underlying predictable and unpredictable evolution
4. determine conditions under which to expect various degrees of predictability
5. establish role of predictable evolution in speciation
6. uncover generality of predictable patterns across taxa
With the GMDS approach, a researcher explicitly derives a priori predictions of phenotypic or genetic change based on a specified set of assumptions for a particular system, and then tests the predictions using comparative and/or experimental data. Generalized models can be built from the understanding
of genetics, biochemistry, development, biomechanics, behavior, and ecology. Predictions generated with GMDS will typically take the form of probabilistic predictions, rather than precise evolutionary endpoints--although, the form of predictions will depend on the goals and state of knowledge of the researcher. For example, one might predict phenotypic trajectories across environments, the relative probability of cis-regulatory versus coding mutations observed in particular genes, particular trait values, or specific changes in certain gene regions.
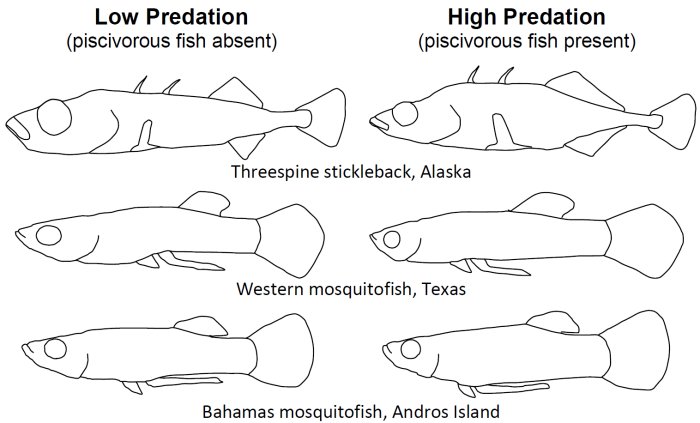
So far, we have explicitly used this approach to formulate and test evolutionary predictions of morphological and locomotor divergence among habitat types in fishes (e.g., see Langerhans 2008, Langerhans and Reznick 2010, Langerhans in press). One of the more thorough tests to date involves body shape divergence between predator regimes, which has been tested in over 20 species from six families across the globe. Remarkably, the vast majority of tests have revealed predictable patterns of divergence.
|